Waseda University team sheds light on self-organization of biological structures
Sat, Nov 23, 2013-
Tags
Researchers at Waseda University in Japan have identified key information to help explain the formation of the “spindle apparatus”, a structure required for cell division. Their findings shed light on the mechanisms behind “self-organization” – an essential characteristic of biological structures.
Organisms are composed of a variety of structures including muscles, internal organs, and brains, all of which are created through a process known as self-organization. In a study published in the online journal, Cell Reports, the research team examined how the spindle apparatus self-organizes. Composed of fibrous molecules called microtubules, this structure is responsible for the segregation of chromosomes between daughter cells.
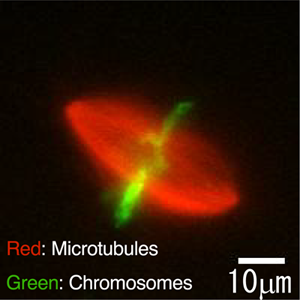
Figure 1: Fluorescent image showing a rugby ball-shaped spindle apparatus inside an egg extract from Xenopus, a genus of aquatic frogs.
Researchers around the world are interested in the mechanisms of spindle formation because if chromosome segregation does not take place correctly in human cells, the process can cause cancer or birth defects. Previous studies have identified molecular motors and a range of other molecules involved in spindle formation. But certain fundamental data remain missing, particularly concerning the relationship between the amount of microtubules and the size and shape of spindles.
Using fluorescence microscopy, Jun Takagi and his colleagues at Waseda University observed self-organizing spindles from the eggs of aquatic frogs. Based on their observations, the team derived a simple mathematical model describing the relationship between the size and shape of the spindle apparatus and the density and amount of microtubules. This successful characterization of the key parameters that determine spindle structures during self-organization is particularly useful in understanding the physical mechanisms of ‘self-organization’ in orderly structures.
Background and purpose of the study
Organisms are composed of a variety of structures including muscles, internal organs, and brains, all of which are created through a process known as self-organization. In the present study, our focus was on the spindle apparatus, a structure responsible for the segregation of chromosomes between daughter cells. The spindle apparatus is formed by self-organization, through molecular-motor assisted assemblage and orientation of fibrous polymers called microtubules. Many researchers around the world have been studying the mechanisms of spindle formation because, if chromosome segregation does not take place correctly, it can cause cancer or birth defects. Previous studies have identified molecular motors and a range of other molecules involved in spindle formation. Certain fundamental data remain missing, however, particularly on the relationship among the amount of microtubules and the size and shape of spindles. Our aim was to investigate the quantitative relationship among those parameters by using physical techniques.
Techniques developed during the study
Spindle apparatus assumes a 3D structure and, in metaphase, it looks like a rugby ball (see Figure 1 above) . While most previous studies used epifluorescence microscopy to observe spindles two-dimensionally, we employed 3D observation using confocal fluorescence microscopy in order to determine the size and shape of spindles more accurately. Thanks to the 3D observation approach, we were able to perform quantitative analysis of 3D asymmetry in deformed spindles. Additionally, fluorescence labeling of microtubules allowed accurate measurement of spindle volume and the amount and density of microtubules in each spindle.
We also established a technique for cutting a spindle into two halves using glass microneedles(<1 µm in tip diameter) (Figure 2) . It is generally impossible to directly manipulate spindles under microscope, because spindles are contained in cells. Fortunately, spindles (30 to 50 µm in length)that self-organized in Xenopus egg extracts, as used in our study, have no cell membranes around them, permitting direct manipulation. A physical technique such as cutting was desirable, as it did not affect properties of the media surrounding the spindles during direct manipulations of spindle size and shape besides microtubule amount and density.
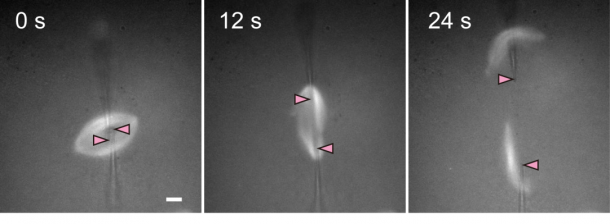
Figure 2: Cutting of a spindle. Spindle microtubules were labeled with a fluorescent dye. To complete the cutting, a pair of glass microneedles were inserted into the spindle and moved to the opposite directions. Needle tip positions are shown by pink arrowheads. The scale bar indicates 10 µm.
Results and conclusions drawn from the study
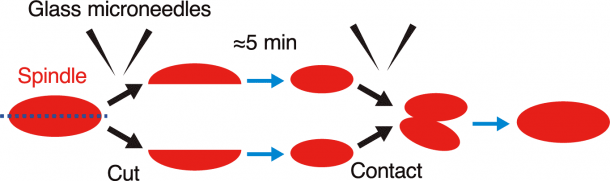
Figure 3: Schematic diagram of the experimental results. When two fragments of a cut spindle come into contact with each other, they fuse together.
Our 3D observation of metaphase spindles that self-organized in Xenopus egg extracts revealed that spindle shape and microtubule density were constant irrespective of spindle size, whereas spindle size was correlated with the microtubule amount. We quantitatively defined the spindle shape and the microtubule density, based on which we successfully derived a simple equation describing the relationship among all the parameters. In this equation, spindle size is explained by microtubule amount in addition to the other parameters that are independent of spindle size (i.e., the spindle shape and microtubule density) .
When a spindle was cut into two fragments using glass microneedles, each fragment regained its original spindle shape and microtubule density within five minutes of cutting (Figure 3) . This indicates that the independent associations between spindle size and spindle shape or microtubule density was maintained even in the cut fragments. Regarding the microtubule amount in each fragment, it was reduced by half or more due to cutting and, at the same time, each fragment became smaller than the original spindle. These findings again indicate preservation of the correlation between spindle size and microtubule amount in the cut fragments. Furthermore, when two cut fragments were allowed to contact each other, they fused together and eventually became a single spindle resembling the one before cutting (Figure 3) .
These results demonstrate that spindle size is correlated with microtubule amount, and that spindle shape and microtubule density are dynamically maintained and unaffected by the physical intervention of ‘cutting.’
Potentially universal effects and social significance of the study
Our successful characterization of the key parameters that determine spindle structures during self-organization is particularly useful in understanding the physical mechanisms of ‘self-organization’ in orderly structures. It is also expected to help elucidate how chromosome segregation is precisely achieved, which is of great importance from a biological as well as medical viewpoint. We believe our findings give clues to the mechanisms of self-organization seen in various other biological structures and, eventually, will facilitate designing of artificial structures utilizing biological materials.
The study reported here was conducted with the financial support of a Grant-in-Aid for Scientific Research. The results have been published in an article entitled ‘Using micromanipulation to analyze control of vertebrate meiotic spindle size’ in Cell Reports, an online journal from Cell Press.
Authors
- Jun Takagi, Research Associate, Faculty of Science and Engineering, Waseda University
- Takeshi Itabashi, Lecturer, Faculty of Science and Engineering, Waseda University
- Kazuya Suzuki, Ph.D. Candidate, Faculty of Science and Engineering, Waseda University; JSPS(Japan Society for the Promotion of Science) Research Fellow (DC1)
- Tarun M. Kapoor, Professor, Rockefeller University
- Yuta Shimamoto, Postdoctoral Fellow, Rockefeller University
- Shin’ichi Ishiwata, Professor, Faculty of Science and Engineering, Waseda University; and Director, Waseda Bioscience Research Institute in Singapore (WABIOS)
Links
Research paper on Cell Reports













