KAKUI Yasutaka, Assistant Professor
Changes in chromatin architecture during the cell cycle
Human genetic information (the human genome), when stored in linear DNA, is about two meters long. In eukaryotic cells, DNA exists as chromatin, a string-like structure connecting a number of bead-shaped units (nucleosomes) wrapped around complexes of a protein called histone (Fig. 1).
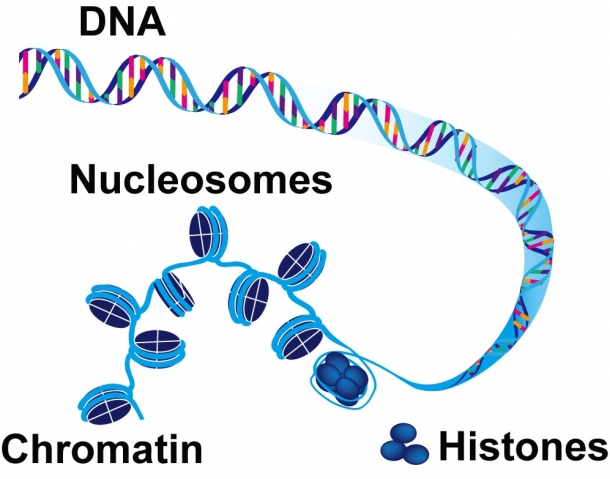
Fig. 1 About 150 base-pairs of DNA wrap around a histone protein complex, connecting to form chromatin.
The series of processes involved in cell division, in which one cell divides into two cells for proliferation, is called the cell cycle. The cell cycle consists of interphase, in which cells perform regular activities and DNA replication, and mitosis, in which the DNA and cellular components are divided into two; the structure of chromatin changes significantly between interphase and mitosis. In the interphase, chromatin is packed in the nucleus like spaghetti in the bowl, while in mitosis, chromatin forms chromosomes, which can be observed with an optical microscope. I have been pursuing research to elucidate how these structural changes in chromatin during the cell cycle are controlled at the molecular level.
Elucidating structural changes in chromatin by observing DNA sequences
Condensin is a ring-shaped protein complex whose primary function is to condense chromatin to form chromosomes during mitosis. Condensin bundles chromatin by connecting it at various points. When chromatin condenses in mitosis, there should be a change in the way it is bundled by condensin, and I wanted to capture that process (Fig. 2).
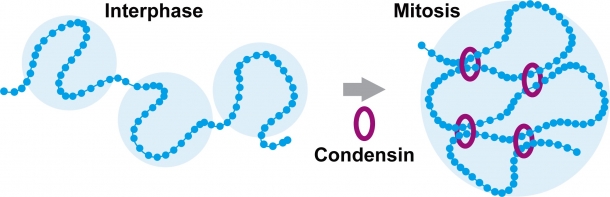
Fig. 2 Condensin and other factors are involved in changes in the spatial arrangement of chromatin during interphase and mitosis.
The main way of studying chromatin structure is the observation using a fluorescence microscope or an electron microscope, but that does not provide sufficient information to reconstruct the internal structure of chromosomes. Therefore, we decided to determine which parts of DNA are close to each other based on DNA sequence information using the high-throughput chromosome conformation capture (Hi-C) method, and investigate their relationship in three-dimensional space. We fixed the cells and fragmented the DNA while maintaining the spatial conformation of chromatin structure, then connected the DNA fragments. This allowed us to capture DNA regions that are in close proximity to each other inside the DNA contiguous domains. Then, by comprehensively reading the sequence of the DNA contiguous domains using a high throughput next-generation sequencer, and comparing them to the original DNA sequence, we achieved determination of the spatial arrangement of chromatin.
For the analysis we chose fission yeast because its genome is only about 1/200 of the size of the human genome and has only three chromosomes. The features of fission yeast genome make high-resolution analysis possible. Our analysis revealed that during the process of chromatin condensation to form mitotic chromosomes, the interactions between different chromosomes or the arms of one chromosome decrease, while the interactions within one arm, in places that are close to each other, increase (Fig. 3). By means of other analyses and experiments, we were able to elucidate in detail the process by which condensin condenses chromatin.
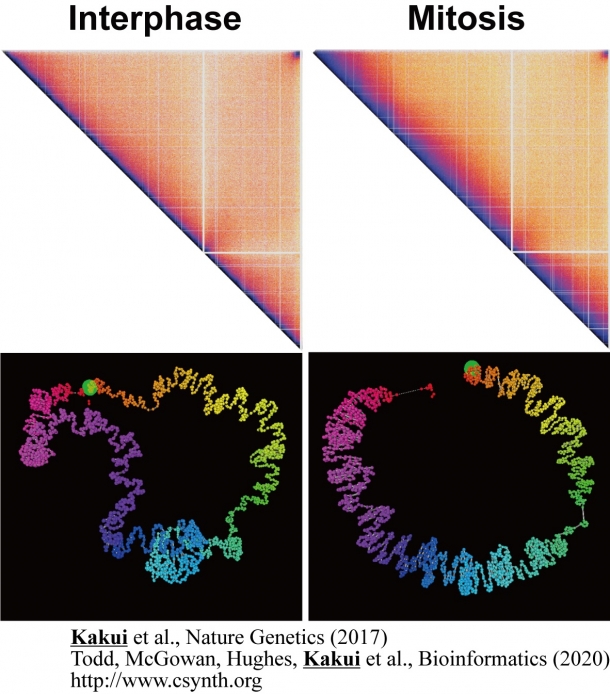
Fig. 3 Hi-C interaction maps and reconstructed chromatin organization in interphase and mitosis. During mitosis, chromatin condenses due to strong medium-range interactions.
Aiming to elucidate the molecular mechanism of abnormalities in meiosis
Among the various kinds of cell division, I am particularly interested in meiosis, which produces gametes (germ cells, oocytes and sperm). Since meiosis involves more complex changes than the division of somatic cells (non-germ cells), understanding the behavior of chromatin during meiosis is a major challenge.
Currently, I am studying a structure called chiasma, which connects homologous paternal and maternal chromosomes in the first meiotic division. Chiasmata are the sites of chromosome crossing over due to recombination; this is important for creating genetic diversity in gametes, but we do not yet know how the place where chiasmata can occur is determined. Therefore, I would like to create an experimental system in which recombination only occurs in one place, so that I can elucidate the molecular mechanism of chiasma formation.
I believe that elucidating the molecular mechanism of meiosis will be an important contribution to reproductive medicine, which is gaining increasing attention in society. When abnormalities occur in meiosis, chromosome distribution is not normal either; this can cause infertility, miscarriages, and congenital anomalies. It is known that these abnormalities are more common at advanced maternal age, but if we can determine what kind of abnormalities are present in the structure of chromatin in aged eggs, in the future it may be possible to diagnose abnormalities in an egg before introducing sperm.
Simply being impressed was my starting point
I set out on the path to becoming a researcher when I was fascinated by the structure of chromosomes in a book of science materials in elementary school. From that experience, I would like to convey the fun of science in the way that even elementary and junior high school students can understand. For example, a number of studies have shown that the mechanism of chromatin condensation to form chromosomes is an elegant approach for determining the accurate distribution of long DNA in post-mitotic cells that are micrometers in size. I would like to unravel that simple but complicated mechanism as a way of giving back to society.
Interview and composition: Shinya Nakazawa
In cooperation with: Waseda University Graduate School of Political Science J-School










